The escalating prevalence of antimicrobial resistance (AMR) in skin bacteria, particularly Cutibacterium acnes and Staphylococcus epidermidis, has become a significant global clinical concern due to the widespread use of antibiotics in treating acne vulgaris. While high resistance rates are well-documented in acne patients, there is a growing need to understand the resistance status of the general population’s skin microbiome to determine if these resistant strains are establishing themselves independently of direct treatment. This study was conducted to investigate the baseline AMR status in individuals with no history of acne therapy, serving as a vital “solution” to clarify whether the observed resistance in clinical settings is primarily driven by specific antimicrobial interventions or reflects a broader environmental shift in commensal bacteria.
Methods
Researchers analyzed skin swabs from 119 healthy university volunteers across two periods (2014 and 2019) who had no history of antimicrobial acne therapy. They identified S. epidermidis and C. acnes isolates, evaluating their antimicrobial susceptibility and resistance determinants through PCR and DNA sequencing. These results were statistically compared against isolates previously collected from acne patients to determine the impact of clinical antibiotic use on the commensal skin microbiome.
Key Findings
- Presence of Baseline Resistance: Macrolide-resistant S. epidermidis and C. acnes were isolated from healthy volunteers despite their lack of prior exposure to acne-specific antibiotic treatments.
- Significantly Lower Rates in Volunteers: In 2019, resistance rates for roxithromycin and clindamycin in volunteers were 4.1% and 2.0% for C. acnes, respectively, which is significantly lower than the rates found in acne patients.
- Distinct Resistance Mechanisms: The primary resistance factor for macrolides in S. epidermidis was the erm(C) gene, whereas C. acnes resistance was predominantly linked to 23S rRNA mutations.
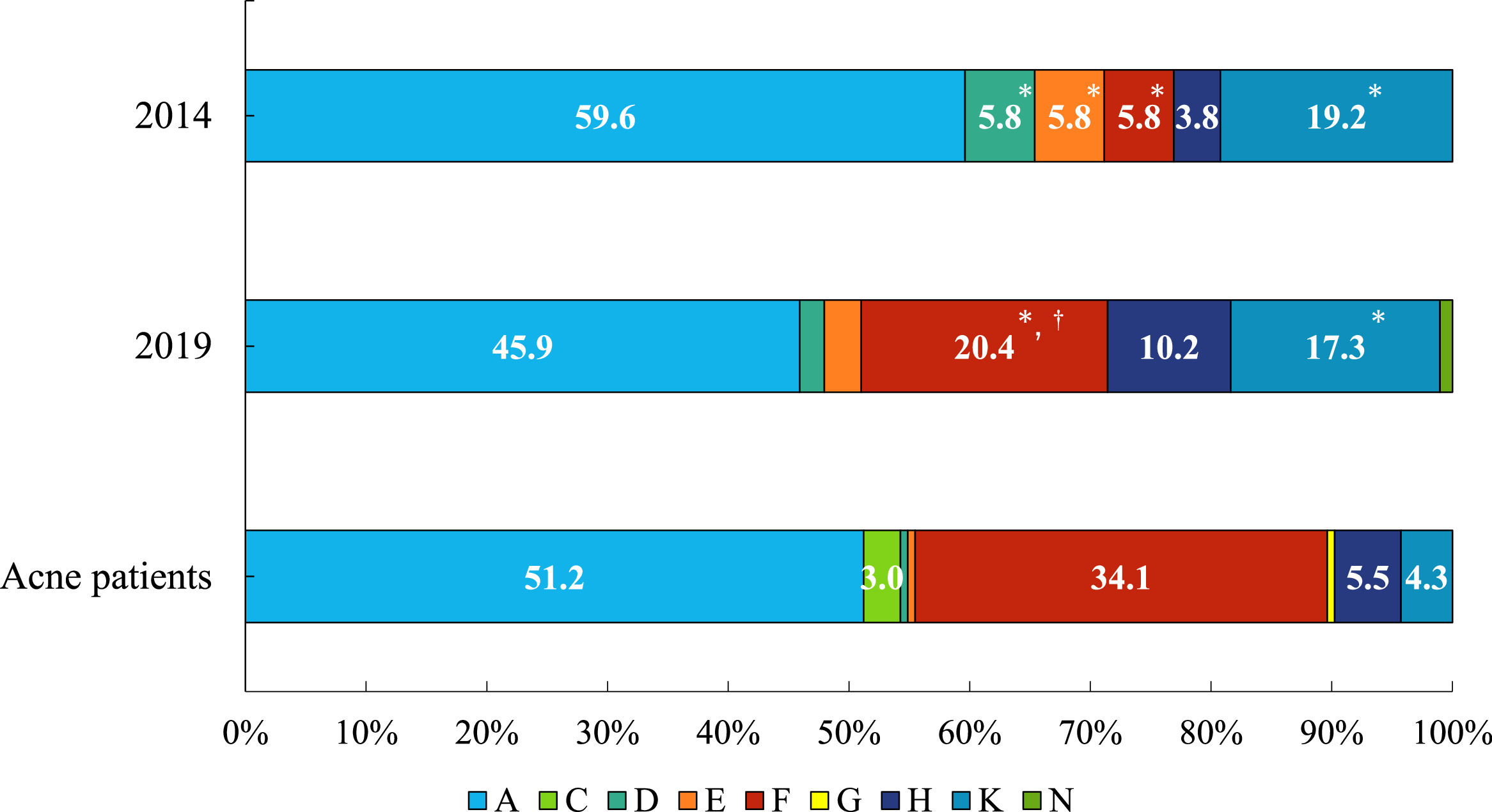
- Clade Distribution Variations: Healthy volunteers exhibited a significantly higher population of clade K (associated with skin health) and a lower population of clade F compared to acne patients.
- Emerging Trends: There was a statistically significant increase in the proportion of the more virulent and resistant-associated clade F among healthy volunteers between 2014 and 2019.
The novelty of this research lies in its comparative demonstration that while antimicrobial-resistant strains are present in the general population, the intensive use of antimicrobials for acne treatment is a substantial driver that significantly amplifies AMR rates in skin flora. These findings have critical future implications, emphasizing the urgent need for the judicious use of antimicrobial agents in dermatology to preserve the diversity of the skin microbiome and prevent the further emergence of resistant opportunistic pathogens. Furthermore, the study suggests that monitoring the microbiome of untreated individuals is essential for a comprehensive global strategy against antimicrobial resistance.
Link to the study: https://www.sciencedirect.com/science/article/pii/S2213716526000494
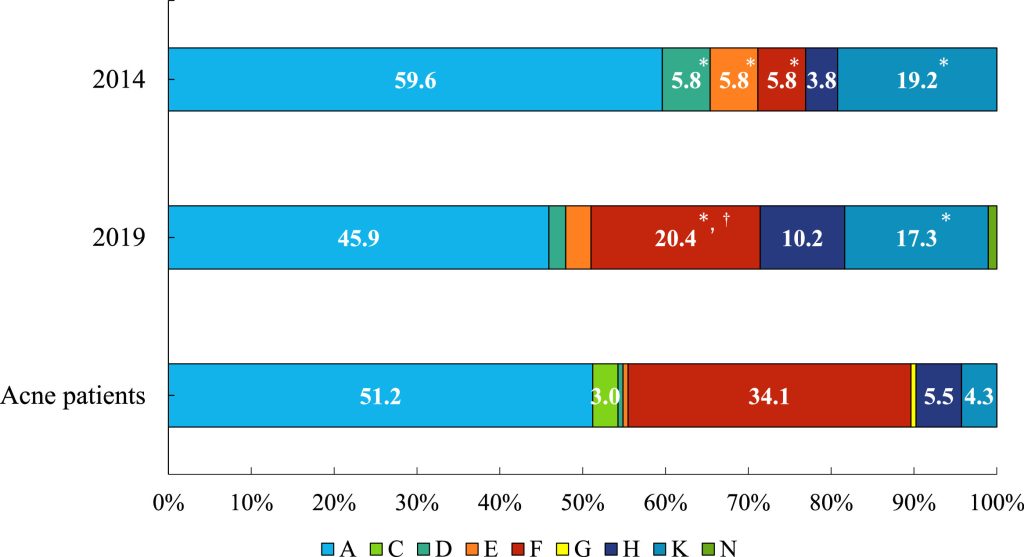
Figure: The distribution of Cutibacterium acnes single-locus sequence typing (SLST) clades is shown among isolates from healthy volunteers in 2014 and 2019, as well as among those from patients with acne. The stacked bar charts illustrate the proportions of SLST clades among C. acnes isolates obtained from the three groups. 1) Isolates from healthy university student volunteers in 2014 (n = 52). 2) Isolates from healthy university student volunteers in 2019 (n = 98). 3) Isolates from patients with acne (n = 164). Clade A was the most prevalent in all three groups. Clade F was significantly more prevalent in isolates from patients with acne than in isolates from healthy volunteers in both 2014 and 2019. Conversely, clade K was more prevalent in isolates from healthy volunteers than in isolates from patients with acne. * significantly different compared with isolates from acne patients (healthy volunteers in 2014 compared with acne patients, or healthy volunteers in 2019 compared with acne patients); † significantly different between 2014 and 2019 (Fisher’s exact test, P < 0.05).